In this blog series, we are exploring the role of key genes that contribute to the pathophysiology of myeloid malignancies, a group of haematopoietic clonal disorders.
Myeloid malignancies are influenced by various genetic and epigenetic influences, manifested by excessive proliferation, abnormal self-renewal, and/or differentiation defects within haematopoietic stem cell (HSC) and myeloid progenitor cell populations. In our first blog, we explored the function of the FLT3 gene and how various alterations to this gene can prompt excessive cell proliferation leading to acute myeloid leukaemia (AML), and other myeloid malignancies.
This time, we will cast a spotlight upon a gene with an altogether distinct function from FLT3, that can nevertheless have a profound effect upon cells of haematopoietic lineage when abnormalities arise. The gene in question is known as KMT2A, a protein-coding gene located within human chromosome 11
KMT2A (Lysine methyltransferase 2A), can be considered part of the epigenetic machinery, and encodes a histone-modifying enzyme called lysine methyltransferase (KMT). This enzyme is responsible for the regulation of chromatin via the covalent modification of histone proteins with the addition of methyl groups1. The methylation of histone proteins can have a significant influence upon chromatin state, and regulates the accessibility of DNA to factors that mediate transcription, DNA repair, and DNA replication.
KMT enzymes are known as writers: enzymes that catalyse the addition of chemical groups to histone tails or to DNA2, and make up part of the epigenetic machinery, which consists of:
Since all three components of the epigenetic machinery are key in gene regulation and chromatin state (whether a transcription is activated or repressed), any abnormalities in genes encoding these components can give rise to an altered chromatin conformation and incorrect gene expression. As such, KMT2A mutations have been implicated in numerous genetic malignancies.
In myeloid malignancies, recurring genetic abnormalities are extremely common, and these can provide important prognostic information and guide therapeutic decisions.
KMT2A rearrangements have been widely reported in a number of myeloid malignancies. The most common alterations involving KMT2A are mutations (3.62%), fusions (0.13%), losses (0.10%), amplifications (0.07%)2. Of particular interest in myeloid malignancies though, are rarer KMT2A aberrations known as partial tandem duplications (PTD). KMT2A-PTDs are characterised by a large internal duplication spanning 6-8 exons, and has been reported in up to 10% of AML and myelodysplastic syndromes (MDS). This recurrent mutation is associated with adverse risk3, with high KMT2A-PTD RNA levels correlating with poor patient prognosis and survival rates.
Despite the high prevalence of KMT2A fusions and known associated outcomes (KMT2A has at least 80 identified fusion partners), the genomic mechanisms that regulate KMT2A-PTD expression are not yet fully understood, and this is likely due to difficulty in identifying KMT2A-PTDs by traditional cytogenetic means. The better characterised KMT2A fusion mutations however, have been shown to deregulate the transcriptional program of cells. In turn, this drives the promotion of specific gene expression patterns that include high expression of HOX genes which are known to contribute to leukaemogenesis4.
KMT2A-PTDs are somewhat cryptic mutations that prove problematic to routinely detect. Conventional cytogenetic approaches such as karyotyping do not normally detect KMT2A-PTDs and the commonly repeated exon regions are too small for detection by FISH. Therefore, the most frequently used method of detection of KMT2A-PTDs is long-range RT-PCR using DNA extracted from a patient’s bone marrow.
However, since most myeloid malignancies are widely heterogenic, running multiple PCRs for each candidate aberration is laborious and costly. It would be preferable to integrate KMT2A-PTD testing into a multiplex assay with the ability to interrogate a patient’s genome for multiple relevant mutations simultaneously. For this reason NGS technologies represent a more useful tool for detecting somatic mutations linked to AML and other myeloid malignancies. However KMT2A-PTDs continue to cause issues here as they are too large to be reliably detected by amplicon-based NGS.
Fortunately, hybridisation-based NGS succeeds where others fail—and has been shown to detect KMT2A-PTDs with a greater sensitivity versus PCR5. By applying a capture-based NGS protocol with probes covering the entire KMT2A region, the presence of a PTD can be reliably interrogated by obtaining the ratio of commonly repeated exon regions in KMT2A-PTDs, against non-duplicated regions. Using the hybrid capture approach, KMT2A-PTDs can now be detected with exceptional accuracy and reproducibility.
This means that myeloid-specific research panels—such as the SureSeq ™ Pan-Myeloid research panel —can offer an affordable and comprehensive platform for the reliable detection of key aberrations associated with myeloid malignancies.
The benefits of hybridisation capture in combination with wide coverage of myeloid-specific aberrations including KMT2A-PTD and other variants, enables accurate detection with high sensitivity—while reducing analysis complexity, assay duration, and costs.

Is NGS set to become the new gold standard in constitutional genetics? Discover how NGS offers greater insight into indels, CNVs, LOH, mosaicism and more.
Read
Designing in situ probes for hybridisation assays can be a tricky, time-consuming task. Read our blog to discover five tips for effective probe design!
Read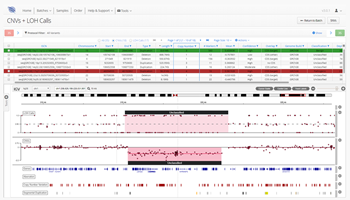
As one of the first steps in many NGS data analysis pipelines, accurate variant calling is often critical to downstream analysis and interpretation. Here, we take a look at variant calling best practice through a modern lens.
Read
A high-quality sequencing library is the linchpin to generating good sequencing data. We discuss our six top tips to help you improve your sequencing library.
Read
We discuss the development and current state of sequencing technologies, and where the future of NGS may take us...
Read
FISH is a cytogenetic technique utilised in labs to detect chromosomal abnormalities in both cancer and constitutional specimens. In this blog learn about the advantages of FISH...
Read
This blog will discuss FLT3’s normal function, its implications in myeloid malignancies, and the role of NGS in genetic identification and disease management of patients with FLT3 genetic alterations.
Read
While liquid biopsy may present an attractive alternative to a solid biopsy, it also has limitations. Here, we shed light on some advantages, limitations, and future outlook for liquid biopsy in oncology clinical practice.
Read
Find out about the benefits and differentiating factors of the three most commonly used NGS technologies; targeted gene panels, whole-exome sequencing (WES) and whole-genome sequencing (WGS).
Read
Next generation sequencing (NGS) is now in routine use for a broad range of research and clinical applications. Facilitating the detection of a wide variety of mutations, focus has never been higher on the value of making the correct choice for the initial sequence enrichment step, which, if poorly designed, can be a source of bias and error in the downstream sequencing assay.
Read