The study of constitutional genetics is the study of how genetic variation relates to human phenotypes and diseases. Accordingly, constitutional disorders are generally considered innate or inborn disorders which have a strong genetic influence. Next generation sequencing (NGS) has revolutionised our understanding of constitutional genetics, not least by enabling the identification of significant numbers of pathogenic mutations that were previously unknown.
Developmental delay (DD) and intellectual disability (ID), for example, are major forms of neurodevelopmental disorders (NDDs) that have been associated with several types of chromosomal aberrations. But the advent of NGS truly brought to light the significant number of different genetic factors that were linked to NDDs and has proven invaluable to improving the diagnostic process and offering new targets for therapy1.
This blog explores the tools used for identifying the genomic underpinnings of constitutional disorders and how next generation sequencing (NGS) is beginning to change how we interrogate and understand human disease.
Cytogenetic testing is the examination of chromosomes to identify chromosomal abnormalities such as aneuploidy (one or more extra or missing chromosomes) and structural variants.
While karyotyping was one of the first cytogenetic tools developed for the detection of constitutional variants, several techniques have since been developed to help researchers to examine targeted genomic regions in patients with constitutional diseases - most notably, fluorescence in situ hybridisation (FISH) and chromosomal microarray analysis (CMA). Both methods employ fluorescent probes that hybridise to specific, complementary sequences on a chromosome resulting in coloured signals that can be detected using a fluorescent microscope.
Discover more on how FISH works and why it remains the gold standard in many clinical applications
Both FISH and CMA have been tweaked and there now exists a whole buffet of CMA and FISH flavours to choose from. Microarray-based comparative genomic hybridisation (aCGH) and single nucleotide polymorphism (SNP) arrays, for example, are subtypes of CMA that have improved resolution.
Karyotyping, FISH, and array-based technologies are still used today – and in some situations remain the gold standard. But like all technologies, they have limitations. Karyotyping, for instance, has an extremely low resolution of about 5-10 Mb. This means it is not able to detect small re-arrangements or SNPs.
And while FISH and microarrays have much better resolutions than karyotyping – and remain indispensable tools for certain clinical applications – they can’t do everything.
For instance, while aCGH is still regarded as the gold standard for analysis of copy number variation (CNV) in ID and DD research samples, it does not provide a comprehensive view of possible aberrations associated with a given sample.
Similarly, because FISH is a targeted analysis it will only identify chromosomal changes at the specific binding region of the probe. Great for diagnosis or prognosis of diseases associated with well characterised constitutional variants. But when it comes to exploratory research or diseases linked to many, variable potential variants, an alternative, more thorough approach is required.
NGS has been established as a revolutionary technology for the analysis of constitutional diseases. It provides unmatched resolution with the ability to detect SNPs and small insertions and deletions (indels), at the same time as structural variants (SVs).
NGS also provides clinicians and researchers with various platforms to explore constitutional disease, including gene panels, whole exome sequencing (WES), and whole genome sequencing (WGS).
Gene panels examine a predefined set of genes associated with a particular phenotype or disease state. By limiting the number of genes sequenced in each experiment, they offer a cost-effective strategy to detect several variant types in a single assay.
Moreover, by focussing on a limited set of genes, NGS gene panels generally have a short turnaround time, have a low rate of unspecific or incidental findings and are easy to interpret. Therefore, they are often used when a known gene or list of gene mutations have been linked to or may be the cause of the disease. The CytoSure® Constitutional NGS panel, for example, targets over 700 genes delivering CNV analysis down to single exon level, loss of heterozygosity (LOH) as well as SNV and indel detection all in a single assay.
Mutations within the SHANK3 gene, for example, are thought to result in a decrease in synapse function and cell-to-cell communication between neurons and ultimately, contribute to DD or ID2,3. The clever bait design in this assay coupled with a proprietary CNV calling algorithm facilitates uniform sequence coverage and robust detection of even the smallest CNVs including a 51 kb deletion in the SHANK3 gene.
Additionally, gene panels are enabling researchers and clinicians to identify detection within mosaic samples. For instance, the CytoSure Constitutional NGS panel detected a 7 Mb deletion at 15q14-q15.1 associated with unexplained DD within a mosaic sample.
OGT’s expertise in bait design ensures uniform sequencing coverage of the desired regions. This, coupled with a proprietary CNV calling algorithm, allows robust detection of even the smallest CNVs.
Exome sequencing is the sequencing of the coding regions (i.e., exons) of the genome. Although the exome only makes up approximately 1% of the whole genome, 85% of all disease-causing mutations are located here.
Because of its ability to provide information on variants across the entire coding region of a genome, WES is often employed as a test when clinicians require more information. For example, WES may be used when a FISH or targeted gene panel assay, are unable to identify a pathogenic mutation.
WGS produces data for over 90% of the genome and unlike panels and WES, it does not require enrichment prior to sequencing. It also provides the ability to simultaneously and comprehensively detect SNVs and CNVs, as well as complex variants such as balanced/unbalanced structural rearrangements (e.g., translocations, inversions, and insertions) and repeat expansions.
Unlike WES, WGS provides sequence coverage for the non-coding as well as the coding regions of the genome. And, although WES provides sequence coverage for nearly all of the mutations currently recognised to cause Mendelian diseases, an increasing number of disease-associated variants have been found in non-coding regions. Thus, WGS is an important, testing strategy when targeted and exome sequencing have failed to identify the underlying mutation of a disease.
The ability to efficiently query many genes and genetic regions, as well as detect a whole platter of variation types simultaneously is possibly the biggest advantage of NGS. Additionally, NGS can be used in both a targeted and non-targeted way permitting hypothesis-free, discovery-based research that seeks to identify novel causes of disease.
So, should we all set down our cytogenetics tools immediately and switch to NGS?
NGS offers some advantages over more traditional cytogenetics tools, however, like all technologies, it has limitations. For example, WGS may provide you with a wealth of information but it is also relatively expensive, has high turnaround times, large data storage needs, and may produce a large number of variants of uncertain significance.
In addition, handling and analysing the large amounts of data generated by WGS and WES to provide meaningful results can require complex bioinformatic pipelines and specific expertise.
Some suppliers are addressing this bioinformatics bottleneck by developing easy-to-use software that effortlessly translates cumbersome NGS data into meaningful insights. For instance, OGT’s Interpret software makes it simple for researchers to analyse and visualise a wide range of variants and structural aberrations in NGS data.
While all cutlery is designed for eating, you wouldn’t use a spoon to cut steak. Equally, you probably wouldn’t eat soup with a fork. Much like cutlery, it’s just about choosing the right one for your specific application.
In other words, NGS is not here to replace FISH, microarrays, or even karyotyping but it does serve as a complimentary approach to studying constitutional diseases. For instance, it is common for pathology labs to use FISH to get quick, reliable results but to also run NGS to gain a deeper insight into the underlying pathogenic mutations of a disease.
From the CytoSure NGS Constitutional panel to our easy-to use, powerful NGS Interpret software, OGT offers comprehensive NGS solutions designed to support you across your entire NGS workflow.

Designing in situ probes for hybridisation assays can be a tricky, time-consuming task. Read our blog to discover five tips for effective probe design!
Read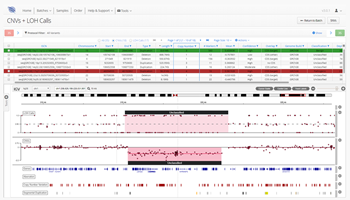
As one of the first steps in many NGS data analysis pipelines, accurate variant calling is often critical to downstream analysis and interpretation. Here, we take a look at variant calling best practice through a modern lens.
Read
A high-quality sequencing library is the linchpin to generating good sequencing data. We discuss our six top tips to help you improve your sequencing library.
Read
We discuss the development and current state of sequencing technologies, and where the future of NGS may take us...
Read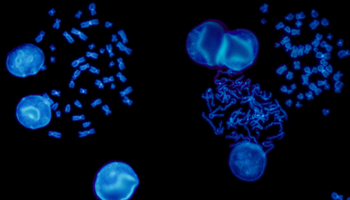
FISH is a cytogenetic technique utilised in labs to detect chromosomal abnormalities in both cancer and constitutional specimens. In this blog learn about the advantages of FISH...
Read
This blog will discuss FLT3’s normal function, its implications in myeloid malignancies, and the role of NGS in genetic identification and disease management of patients with FLT3 genetic alterations.
Read
While liquid biopsy may present an attractive alternative to a solid biopsy, it also has limitations. Here, we shed light on some advantages, limitations, and future outlook for liquid biopsy in oncology clinical practice.
Read
Find out about the benefits and differentiating factors of the three most commonly used NGS technologies; targeted gene panels, whole-exome sequencing (WES) and whole-genome sequencing (WGS).
Read
Next generation sequencing (NGS) is now in routine use for a broad range of research and clinical applications. Facilitating the detection of a wide variety of mutations, focus has never been higher on the value of making the correct choice for the initial sequence enrichment step, which, if poorly designed, can be a source of bias and error in the downstream sequencing assay.
Read