For accurate detection of all types of genetic aberrations, various technologies are required. By combining information from multiple technologies, researchers can analyse complex samples and get the most complete overview of disease-driving mutations.
OGT offers an integrated portfolio of products providing clinical researchers with the most advanced tools available to study hematological disorders, including CytoCell® fluorescence in-situ hybridization (FISH) probes and myProbes® custom FISH probes, SureSeq™ next generation sequencing (NGS) panels and myPanel™ custom panels, and CytoSure® array products and custom arrays.
Our products are backed by deep technical expertise and dedicated customer support.
Developed in partnership with myeloid cancer experts to the latest WHO guidance, the panel can identify novel fusion partners — providing more comprehensive and informative analyses than possible using PCR-driven methods. The figure (above) shows the consistent and confident detection of MECOM overexpression in [A] serial dilutions of HNT-34 cell line as well as [B] research and commercial samples, including positive and negative controls.
View ProductMyeloid malignancies are a heterogeneous group of diseases, associated with a wide range of variants ranging from mutations to structural variations. The panel accurately detects SNVs and indels in genes such as CEBPA, JAK2, CALR and MPL, as well as structural variants including FLT3-ITDs and KMT2A-PTDs, and is able to detect low-frequency SNVs and indels with confidence. The figure (above) shows a PTD detection spanning exons 2-8 of the KMT2A gene.
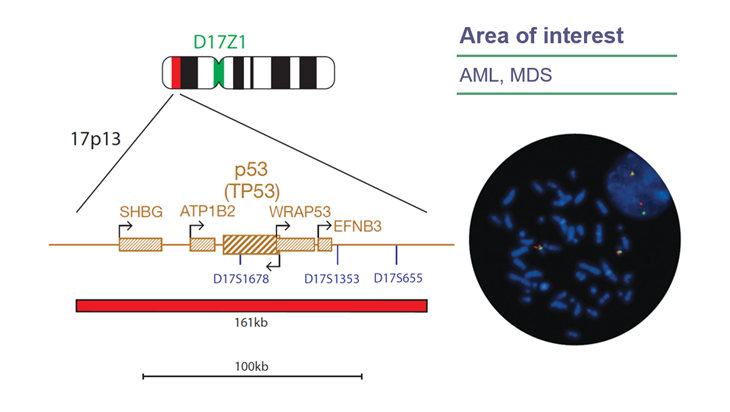
View ProductThe P53 (TP53) Deletion FISH Probe Kit consists of a 161kb probe, labeled in Texas Red, covering the whole P53 (TP53) gene, extending 74kb telomeric to the gene and covering a region centromeric to the gene, to just beyond the marker D17S655; and a probe, labelled in FITC green, covering the chromosome 17 centromere (D17Z1) region.
View ProductThe MLL (KMT2A) Breakapart FISH Probe Kit consists of an 87kb probe, labeled in Texas Red, covering a region telemetric to the MLL (KMT2A) gene including the marker SHGC-111513 and a FITC green probe covering a 170kb region centromeric to the MLL (KMT2A) gene spanning the CD3G and UBE4A genes.
View ProductOGT’s range of CytoSure® Cancer +SNP arrays combine long-oligo probes for superior copy number variant (CNV) detection alongside single nucleotide polymorphism (SNP) probes – which function using OGT proprietary technology – for accurate identification of loss-of-heterozygosity (LOH). The figure (above) shows a schematic of OGT’s unique SNP probe technology, allowing the use of any reference sample with no restriction digest.
View ProductBenefit from OGT’s extensive array design expertise to produce an array matching your precise specifications. These arrays are ideal if you want to know the precise coordinates of an aberration by analyzing specific areas of the genome at high resolution. The figure (above) shows how our custom arrays can be designed in a variety of formats depending on your desired level of focus, with 1, 2, 4, or 8 arrays available per slide to provide the most cost-effective solution for your research.
View Product