Tips include why you should use the appropriate indexes and the low multiplexing guidelines if using a lower (1-8) sample batch for sequencing and what to do if QC yields are lower or higher than expected.
Use the appropriate indexes and the low multiplexing guidelines if using a lower (1-8) sample batch for sequencing. Due to the sequencing chemistry indexes must be sufficiently unique in order to maintain colour balance and facilitate accurate imaging by the sequencer.

Serena Pace
NGS Field Application Scientist, North America
If QC yields are lower than expected check the tape station results. If libraries peak in the optimal area (250-350bp) but have a low yield, they can still be sequenced but may exhibit higher duplication rate and low loading patterns.

Pete Gray
Array and NGS Field Application Specialist, Europe
If QC yields are higher than expected it is likely that the hybridisation and/or wash was not efficient and the captured libraries contain a high amount of off-target DNA. We recommend repeating the hybridisation and wash.

Huiyan Jin
NGS Field Application Scientist, North America
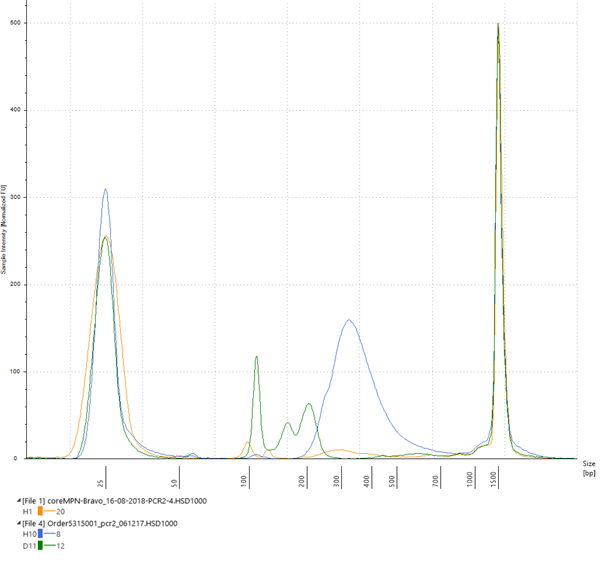
Figure 1: Analysis of amplified captured DNA using an Agilent High Sensitivity D1000 ScreenTape assay. The Blue electropherogram trace shows a peak in the desired size range of approximately 250–350 bp (+/– 10%). The green and yellow traces show examples of low DNA yield and failed PCR2.
If high molecular weight fragments are observed in the TapeStation results, we recommend repeating the hybridisation and wash. The large fragments are indicative of sub-optimal enrichment and over-amplification during PCR.

Kicki Bergefall
FAS Manager – Europe
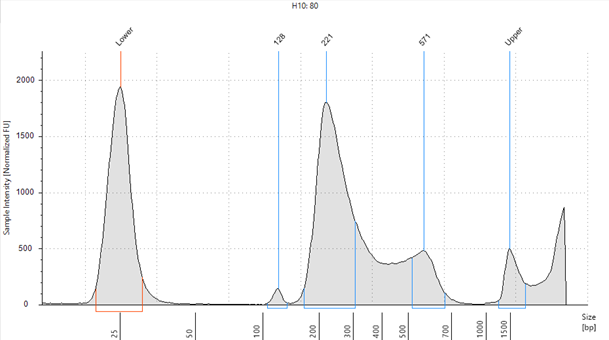
Figure 2: Agilent High Sensitivity D1000 ScreenTape assay showing High Molecular weight fragments at ~500 bp.
