Two broad categories of enrichment assays exist: amplicon (PCR) and hybridisation. As a very general rule, hybridisation-based assays, when designed well, offer superior performance.
Unlike amplicon-based assays, hybridisation is less susceptible to contaminants found in FFPE-derived DNA. Use of an upstream FFPE repair step can significantly improve mean target coverage.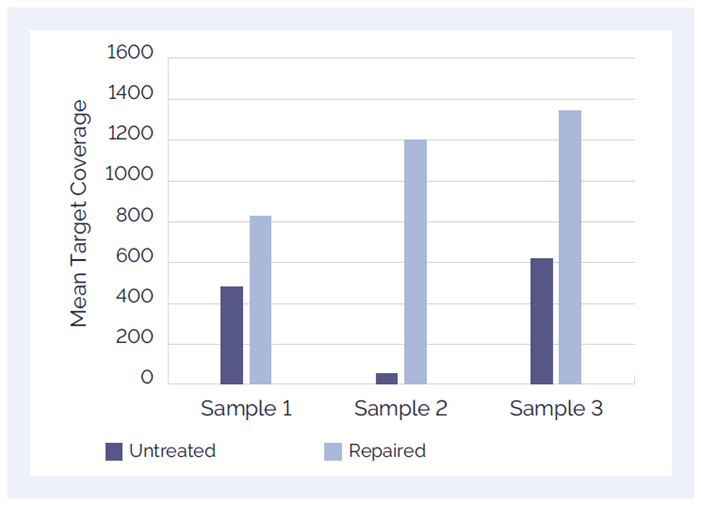
Figure 1. Improved performance with the SureSeq™ FFPE DNA Repair Mix: The SureSeq FFPE DNA Repair Mix significantly improves mean target coverage resulting in more confident calls.
OGT's innovative bait-design delivers uniform and complete coverage of difficult-to-sequence GC rich regions of the genome.
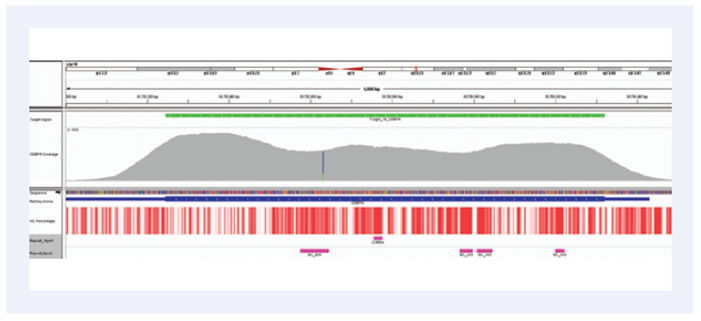 Figure 2. Outstanding coverage uniformity: Illustration of the excellent uniformity of coverage of the CEBPA gene. Depth of coverage per base (gray). Targeted region (green). Gene coding region as defined by RefSeq (blue). GC percentage (red). Repeat regions (pink).
Figure 2. Outstanding coverage uniformity: Illustration of the excellent uniformity of coverage of the CEBPA gene. Depth of coverage per base (gray). Targeted region (green). Gene coding region as defined by RefSeq (blue). GC percentage (red). Repeat regions (pink).
Uniformity of enrichment means that all regions are represented more equally, and that variants present in any region will be called. It also allows much lower average sequencing depths to be used, enabling larger numbers of samples to be multiplexed in a run, and significant cost savings.
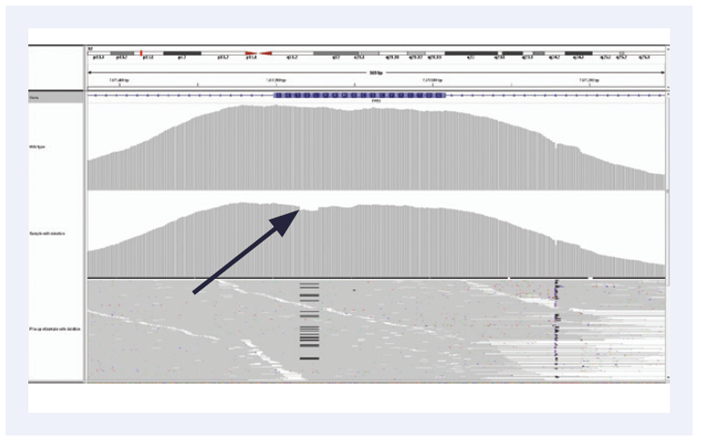 Figure 3. Reliable detection of low frequency mutations: High uniformity of coverage allows the reliable detection of low frequency somatic indels even in FFPE derived DNA. Example is of a 12 bp deletion (c.754_765delCTCACCCATCATC) in exon 3 of TP53, 6% frequency, using the SureSeq Ovarian Cancer Panel.
Figure 3. Reliable detection of low frequency mutations: High uniformity of coverage allows the reliable detection of low frequency somatic indels even in FFPE derived DNA. Example is of a 12 bp deletion (c.754_765delCTCACCCATCATC) in exon 3 of TP53, 6% frequency, using the SureSeq Ovarian Cancer Panel.
Hybridisation-based assays avoid the tendency towards amplification/sequencing bias and error seen with amplicon methods as the degree of multiplexing and number of PCR cycles increases.
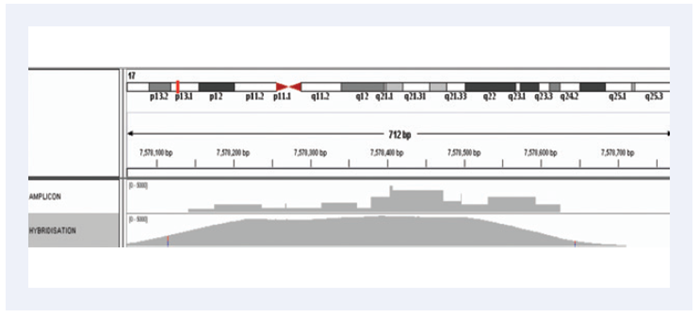 Figure 4. Minimal variation in amplification efficiency: Comparison of amplicon and hybridisation-based enrichment of the GC-rich exons 4 and 5 of the TP53 gene illustrating the superior coverage uniformity.
Figure 4. Minimal variation in amplification efficiency: Comparison of amplicon and hybridisation-based enrichment of the GC-rich exons 4 and 5 of the TP53 gene illustrating the superior coverage uniformity.
Hybridisation has moved on. Well-designed hybridisation assays can now utilise lower amounts of input DNA, whilst still producing clean, bias-free, high quality data.
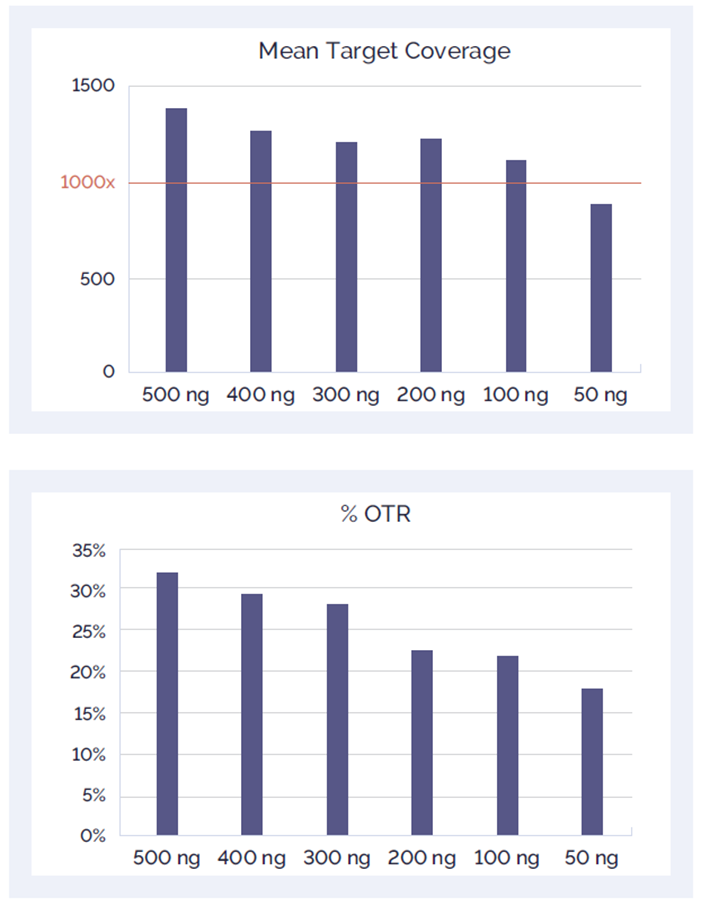
Figure 5. Confident detection with low input DNA: Effect of reduced amount of DNA input on mean target coverage and %OTR.
With a short enzymatic fragmentation step, combined end-repair and adaptor ligation steps, and optimised hybridisation in as little as 30 minutes, you can go from sample to sequencer in a single day.
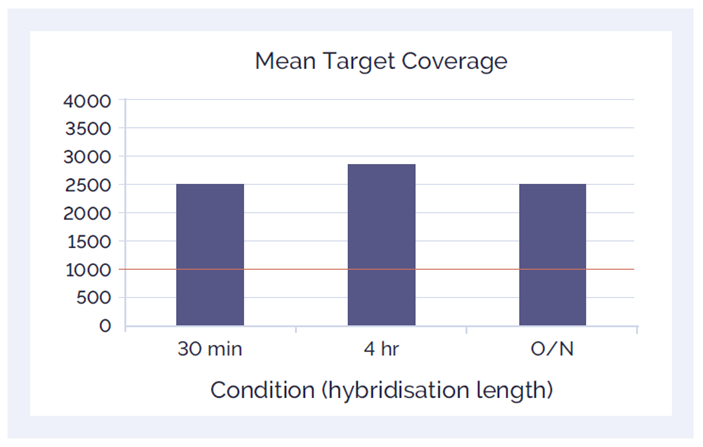 Figure 6. Hybridisation quality - amplicon speed: The hybridisation step has been optimised to take as little as 30 minutes with good quality DNA for the SureSeq Core MPN Panel, without compromising results.
Figure 6. Hybridisation quality - amplicon speed: The hybridisation step has been optimised to take as little as 30 minutes with good quality DNA for the SureSeq Core MPN Panel, without compromising results.
