What is aCGH used for?
Sample and reference DNA are labelled with different fluorescent dyes.
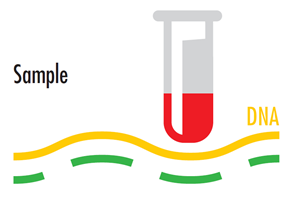
Sample DNA is typically labelled with Cy3 — which looks red under regular light but fluoresces “Green” (575nm) under laser excitation.
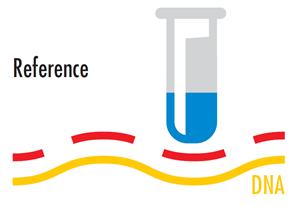
Reference DNA is typically labelled with Cy5 — which looks blue under regular light but fluoresces “red” (675nm) under laser excitation.
Samples are mixed into a single tube.
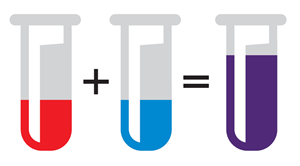
The mixture is pipetted onto a “gasket” slide with chambers which hold the solution.
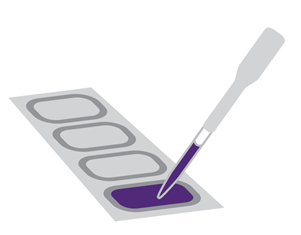
The microarray slide is then held in place against this, ready for hybridisation.
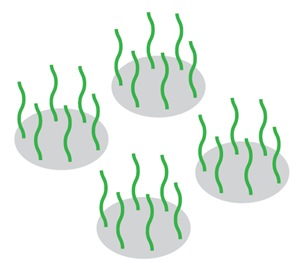

Microarrays contain tightly packed “spots” of DNA oligos, also referred to as probes. Each spot is usually designed to target a different region of the genome.
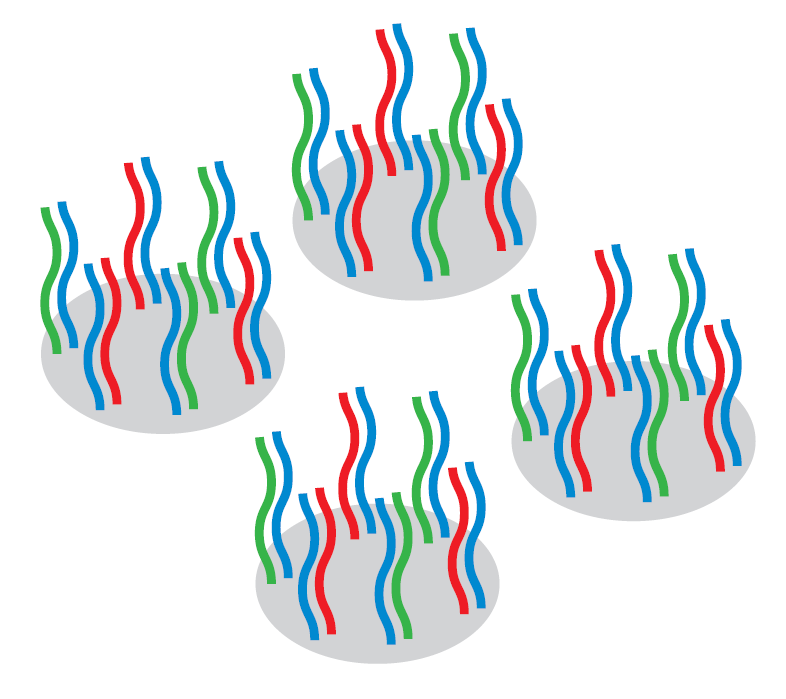

Both the sample and reference labelled DNA compete to bind to the probes, in a process known as “competitive hybridisation.”
Slides are scanned — each spot shows the relative amounts of sample vs reference DNA at a particular genomic sequence.
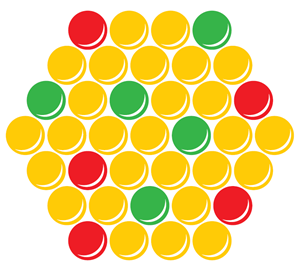

Once scanned, each spot indicates the relative amounts of the sample DNA against the reference DNA at the respective genomic locus. Spots with less sample than reference are indicative of a loss, those with more sample than reference are indicative of a gain.
Purpose-built software allows for analysis and interpretation of aCGH results.
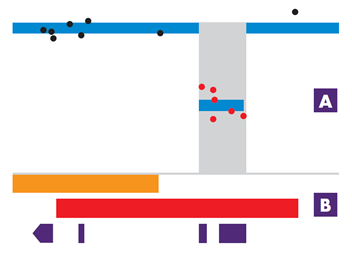
[A] Probe and variant visualisation.
[B] Database tracks.
CytoSure Interpret software uses sophisticated algorithms to define areas of CNV or LOH. Track-based analysis allows for interpretation of these regions against external and internal databases.
