SureSeq myPanel™ is a regularly updated, expert-curated library of pre-optimised cancer panel content for you to select from. Simply mix and match the gene, exonic or intronic content you need to create an NGS cancer panel that meets your exact requirements.
Our unique panel design coupled with hybridisation-based enrichment offers unparalleled coverage completeness and uniformity. This allows the accurate detection of difficult to sequence, low-frequency variants and minimises the requirement for supplementary fill-in with Sanger sequencing.

Detect low-frequency variants consistently with confidence and minimise the requirement for supplementary fill-in with Sanger sequencing

No more laborious in-house optimisation, decreasing assay development time

Sequence only what’s relevant for your cancer research, increase throughput and save on sequencing reagents

Get the most comprehensive insight into disease-driving mutations
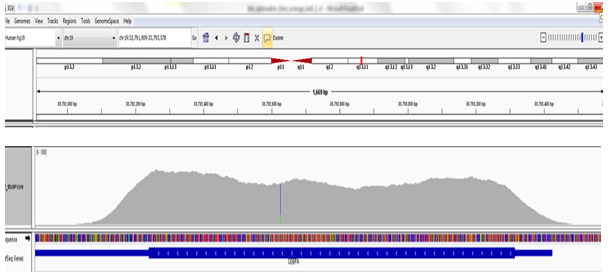
Mutations in the CEBPA gene are among the most common molecular alterations in AML and bear important significance. Sequencing of the CEBPA gene is often hampered by a repetitive nucleotide sequence and a high GC-rich content, which can lead to technical challenges in assay design, requiring a supplementary fill-in with Sanger sequencing. OGT’s expert bait design overcomes these issues, offer a high level of uniformity coverage for a notoriously difficult gene to sequence.
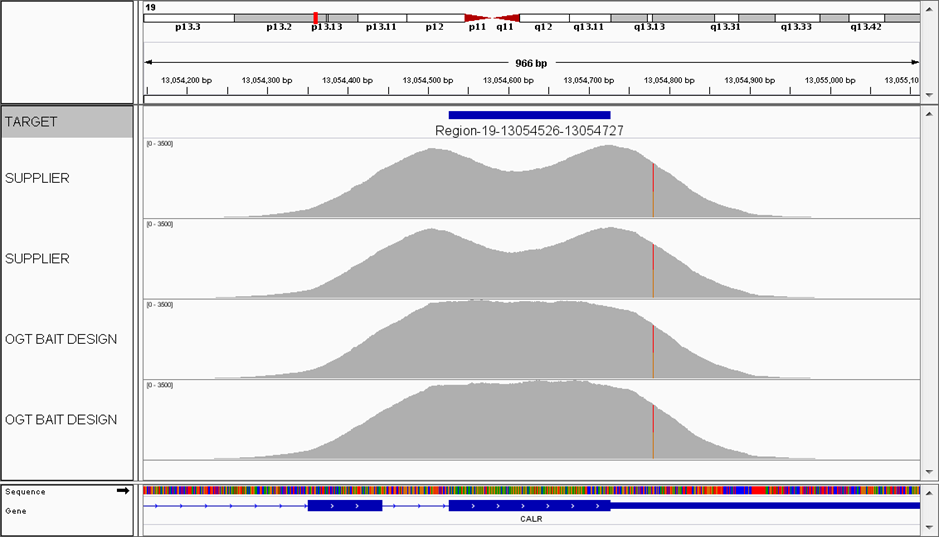
The CALR gene is commonly mutated in various myeloproliferative neoplasms (MPNs). It is critical to identify key CALR indels (types 1 and 2 causing a frameshift) as well as increasingly recognised point mutations in this gene. OGT’s expert bait design offers excellent coverage uniformity across low-complexity regions with low nucleotide diversity (such as CALR exon 9), compared to other commercially available bait designs, which suffer from a considerable dip in coverage.
The BCR-ABL gene fusion is formed following a balanced translocation of chromosome 9 and 22, generating the Philadelphia chromosome, a hallmark of CML. Combine SNV and indel detection with BCR-ABL translocation content and replace multiple assays with a single streamlined NGS workflow for a more comprehensive picture of all your samples.
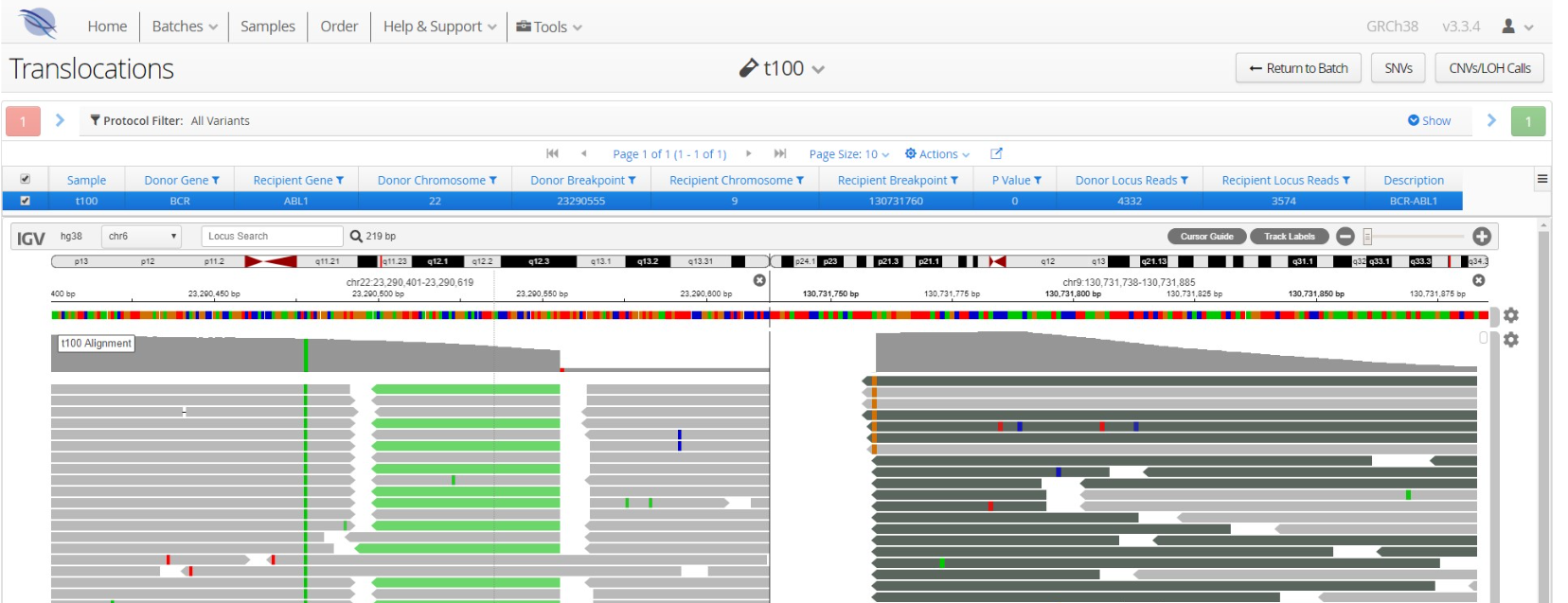
Figure 3: BCR-ABL translocation reported in Interpret. Split-reads covering both BCR (left panel) and ABL1 (right panel) are detected, indicative of the BCR-ABL gene fusion.

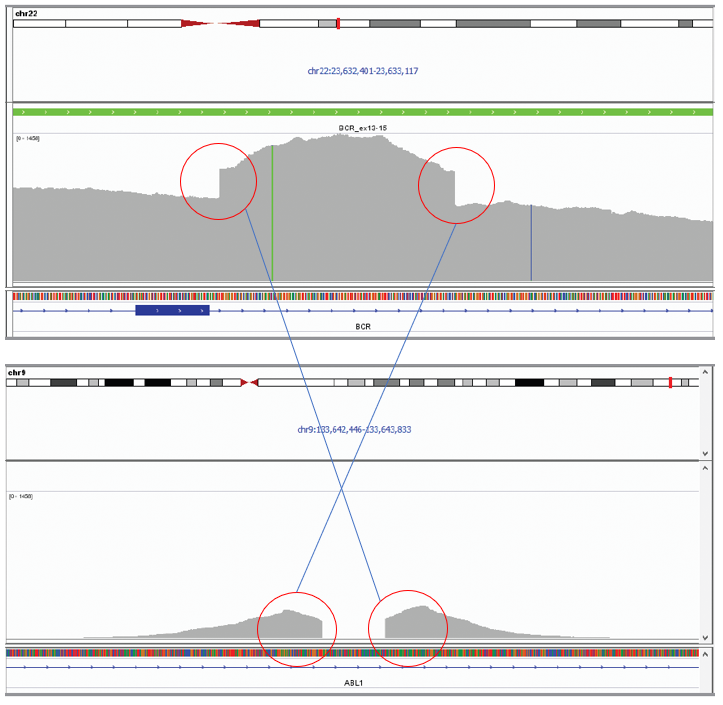
Figure 4: In this example, two translocations are present, with both BCR-ABL1 (chr22:23,632,850/chr9:133,643,198) and ABL1-BCR (chr9:133,643,072/chr22:23,632,613) translocations identified in the CML cell line JK-1. The breakpoints identified by NGS were found to match the coordinates reported in the literature (ref 1).

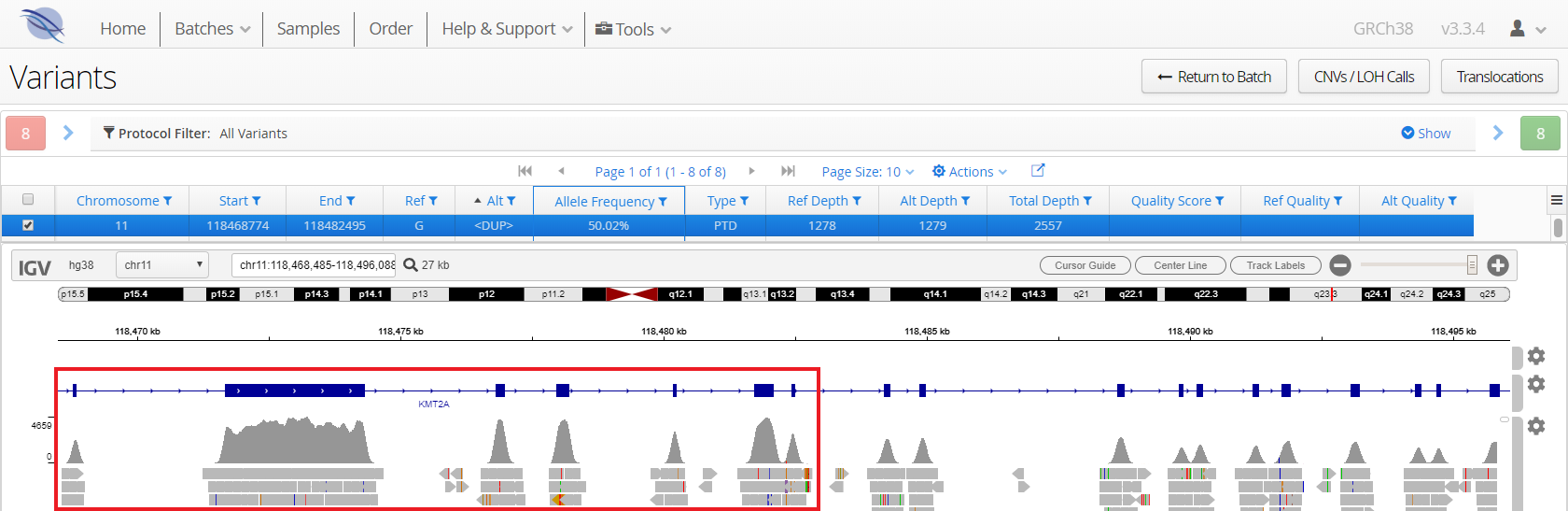
Partial tandem duplications in KMT2A (MLL), frequently observed in AML, are notoriously difficult to detect due to their size, often spanning multiple exons. With OGT’s expertise in hybridisation-based panel design, your SureSeq myPanel offers robust detection of all sizes of KMT2A-PTDs, alleviating the burden of running multiple assays (Figure 5).
* Exon examples not yet available