Myeloid malignancies are a heterogeneous group of diseases, associated with a wide range of variants ranging from mutations to structural variations. The SureSeq™ Myeloid Plus NGS Complete Workflow Solution combines the rapid Universal NGS Workflow Solution V2 hybridisation based target enrichment method together with OGT’s expert bait design to detect 49 key genes implicated in myeloid disorders.
Designed in collaboration with recognised cancer experts, SureSeq Myeloid Plus is a targeted gene panel for next generation sequencing technologies (NGS) to analyse specific mutations in key genes implicated in acute myeloid leukaemia (AML), myeloproliferative neoplasms (MPN), and myelodysplastic syndrome (MDS), providing researchers with a single NGS workflow delivering a comprehensive picture of the genetic make-up of each myeloid sample.
The SureSeq Myeloid Plus Panel accurately detects SNVs and indels in genes such as CEBPA, JAK2, CALR and MPL, as well as structural variants including FLT3-ITDs and KMT2A-PTDs, and is able to detect low-frequency SNVs and indels with confidence.

Detect SNVs and indels in 49 genes implicated in a variety of Myeloid malignancies, together with 44 SNPs as ID markers and 4 sex chromosome genes

Detect low-frequency variants down to 2.5% VAF with confidence

Replace multiple single gene assays with one comprehensive panel

Streamlined library preparation, rapid hybridisation and intuitive software allowing easy variant analysis
Mutations in the CEBPA gene are among the most common molecular alterations in AML, which itself is the most common type of acute leukaemia in adults1,2. Sequencing of CEBPA is challenging due to the presence of repeat regions and the high GC-content of the gene, leading to poor coverage across these regions and potentially missed variants. OGT’s expert bait design overcomes these issues and provides exceptional coverage uniformity, enabling reliable detection of variants and eliminating the requirement for supplementary fill-in with Sanger sequencing (Figure 1).
FLT3 internal tandem duplications (ITDs) are challenging to target, and subsequently detect, because they are by nature repetitive and can be very long. As a result, FLT3-ITDs are generally masked in most panel designs, necessitating additional techniques to obtain a comprehensive genetic picture. OGT employs sophisticated bait designs to generate uniform coverage across, as well as upstream and downstream of the repetitive region. In combination with our complimentary NGS analysis software Interpret, this allows easy detection of FLT3-ITDs ranging from a handful of base pairs to >200 bp (Figure 2).
Other tandem duplications frequently observed in AML are partial tandem duplications (PTDs) in KMT2A (MLL). Similar to ITDs, KMT2A-PTDs are notoriously difficult to detect due to their size, with duplications spanning multiple exons. With OGT’s expertise in hybridisation-based panel design, SureSeq offers robust detection of all sizes of KMT2A-PTDs, alleviating the burden of running multiple assays (Figure 3).
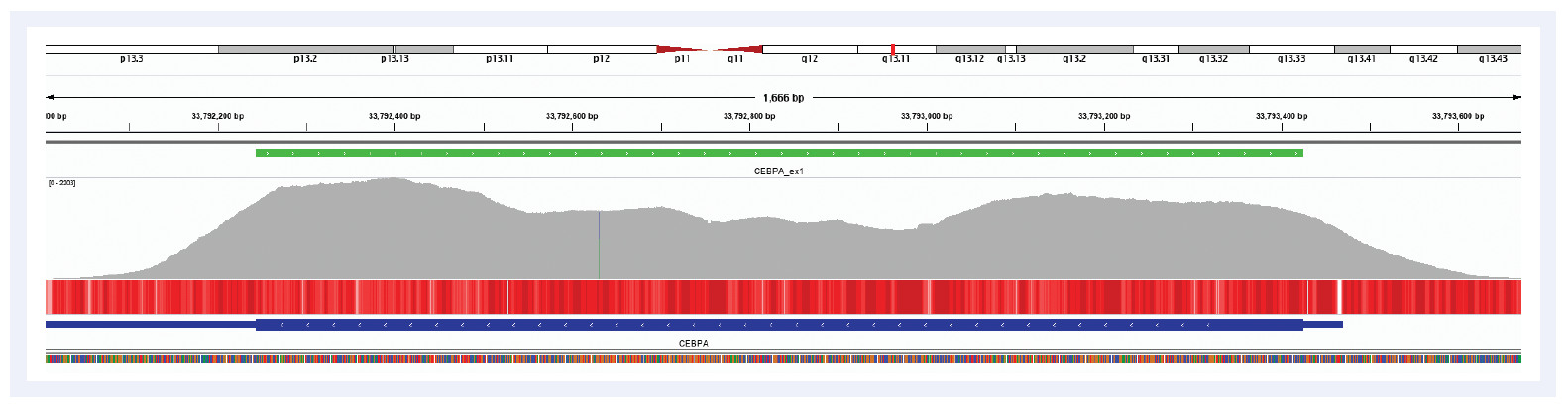
Figure 1: Illustration of the excellent coverage uniformity of the CEBPA gene. Depth of coverage per base (grey). Targeted region (green). Gene coding region as defined by RefSeq (blue). GC percentage (red).

Hybridisation-based enrichment is now well recognised as providing superior results over amplicon-based enrichment technology. The OGT Universal NGS Workflow Solution delivers a straightforward and robust workflow with fewer hands-on steps, and improved turnaround times. For increased convenience and flexibility, the Universal NGS workflow is performed with a combined enzymatic fragmentation, end repair and A tailing step, and convenient bead concentration steps, whilst still delivering libraries of the highest quality. The inclusion of unique Dual Index Adapters increases multiplexing efficiency and confidence, whilst enhancing capabilities to include sensitive applications. The included Universal Hybridisation & Wash Kit simplifies this key step while offering excellent coverage uniformity and reproducibility.
Interpret is OGT’s powerful and easy-to-use NGS analysis solution, facilitating analysis and visualisation of a wide range of somatic variants and structural aberrations. Designed to work seamlessly with all SureSeq panels, Interpret perfectly complements the SureSeq Myeloid Plus Panel, delivering fast and accurate detection of all SNVs, indels, ITDs and PTDs covered by the panel. Following detection, all variants can be easily visualised in the user-friendly variant browser, for an effortless translation of all your myeloid data into meaningful results.
You never have to sequence genes you’re not interested in and can always modify each panel to what’s relevant to your research. If the SureSeq Myeloid Plus Panel doesn’t meet your exact requirements, you can choose from our regularly updated, expert-curated library of pre-optimised cancer content to create your ideal custom SureSeq myPanel™ Myeloid Panel. Alternatively, have a look at the other myeloid panels we have available, including our focused 3-gene SureSeq Core MPN Panel and the SureSeq Pan-Myeloid Panel, incorporating key variants in 70 genes implicated in a wide range of myeloid disorders, or our disease-specific content, such as our SureSeq myPanel NGS Custom AML panels.
The OGT partnership approach is key to providing the highest level of service, working closely with you to understand your unique challenges, customising our approach to meet your exact needs.
* Exon examples not yet available