Chronic lymphocytic leukaemia (CLL) is the most common type of leukaemia in adults. A wide variety of chromosomal abnormalities are associated with CLL, ranging from single nucleotide variants (SNVs) and insertions/deletions (indels) up to large copy-number variations (CNVs), including trisomies.
The SureSeq™ CLL + CNV Panel has been designed in collaboration with recognised cancer experts to detect 13 key genes and 5 chromosomal regions implicated in CLL progression (Table 1). The SureSeq CLL + CNV Panel alleviates the burden of running multiple assays and streamlines your CLL research to deliver a comprehensive genomic profile for each CLL sample using a single workflow.

Detect low-frequency SNVs and indels with confidence

Profile your samples for CNVs in the 5 most commonly aberrant regions in CLL

Replace multiple assays with a single NGS panel, increasing throughput and reducing turnaround time

Analyse your data with Interpret, OGT’s powerful and easy-to-use analysis solution for accurate identification of all variants and CNVs
Investigating both chromosomal aberrations and SNVs/indels is imperative to advance research into CLL progression and treatment. Structural abnormalities are common in CLL and found in more than 80% of CLL cases, the most frequent being del(13q), del(11q), del(17p), del(6q) and trisomy 121. Some of these CNVs cover important tumour suppressors, such as del(17p) resulting in the loss of the TP53 gene. More recently, other genes have also been found to be mutated in CLL, including NOTCH1, SF3B1, MYD88 and BIRC3, adding to the genomic complexity of this leukaemia2.
Due to this genetic heterogeneity, current analysis strategies for CLL require multiple methods to obtain a comprehensive genetic picture, often using microarray or fluorescence in situ hybridisation (FISH) to detect structural abnormalities in combination with NGS for somatic variants. With OGT’s SureSeq CLL + CNV Panel, you can now obtain a more complete understanding of the genetic makeup of CLL progression in each sample using a single assay.
OGT’s expert bait design delivers outstanding uniformity and depth of coverage, offering confident detection of low frequency SNVs and indels down to 1% minor allele frequency (MAF) in 14 genes (Figures 1a & 1b), including SRY and 24 SNPs to allow for easy sample tracking3.
The SureSeq CLL + CNV Panel covers the 5 most common CNVs in CLL enabling reliable detection of these genes/regions. Compared to array data, often considered the gold standard for CNV detection, the events reported with the SureSeq CLL + CNV Panel were 100% concordant, even in genomic regions containing multiple aberrations (Figures 2 - 3). More so, facilitated by OGT’s excellent bait design, loss-of-heterozygosity (LOH) can be identified. With a CNV size detection range from single exon to whole gene, up to complete loss of a chromosomal arm and whole chromosome gains (trisomy 12), your data provides a more comprehensive genetic picture for each sample from a single assay.
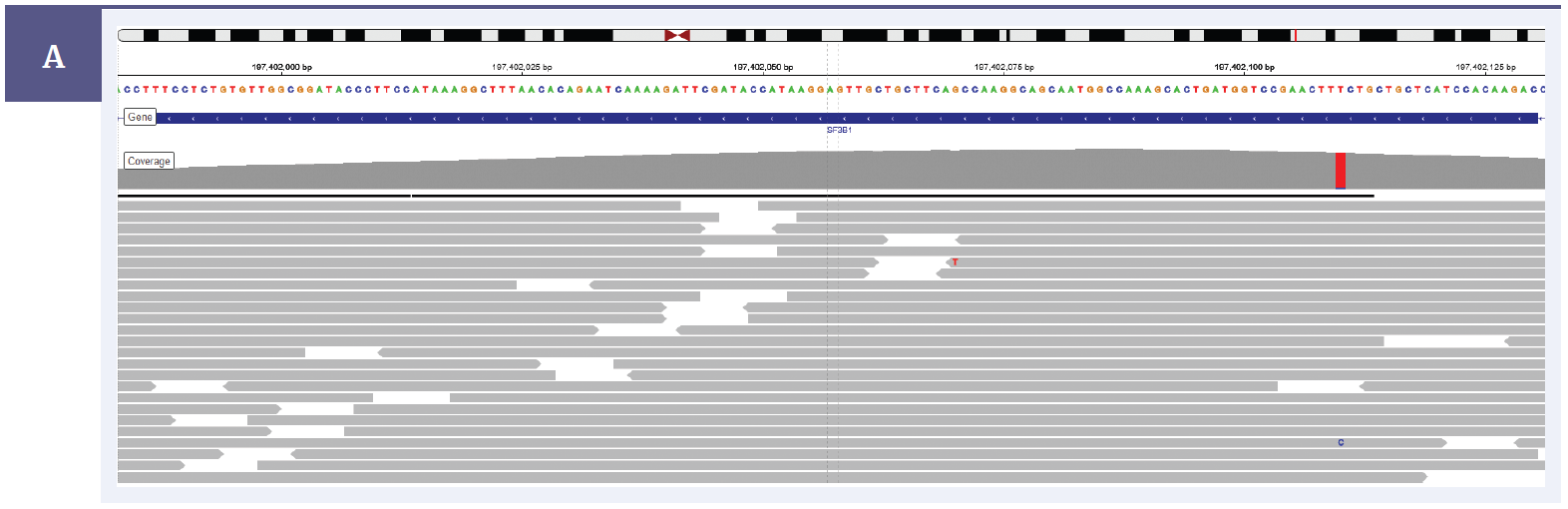
Figure 1a: Illustration of the excellent uniformity and high depth of coverage allowing confident detection of a SF3B1 exon 15 hotspot variant Lys700Glu with 4.8% allele frequency

Interpret is OGT’s powerful and easy-to-use data analysis solution, facilitating analysis and visualisation of a wide range of somatic variants and structural aberrations. Designed to work seamlessly with all SureSeq panels, Interpret perfectly complements the SureSeq CLL + CNV Panel, delivering fast and accurate detection of all SNVs, indels, LOH and CNVs covered by the panel. Following detection, all events can be readily visualised in the user-friendly variant browser, for an effortless translation of all your CLL data into meaningful results (Figure 4).
Does the SureSeq CLL + CNV Panel not meet your exact requirements? (See table 2 for a detailed breakdown of the numbers).
With OGT, you never have to sequence genes you’re not interested in and can always modify each panel to what’s most relevant for your research. Choose from our regularly updated, expert-curated library of pre-optimised cancer content to create your ideal custom SureSeq myPanel™ CLL Panel, or order the SureSeq CLL + CNV Panel right off the shelf.
* Exon examples not yet available

The panel offers comprehensive analysis of genes implicated in CLL pathogenesis, along with examination of critical chromosomal regions associated with the disease, and is an invaluable tool in CLL clinical research.

Anna Sobczyńska-Konefał
Head of the Department of Hemato-Oncology Diagnostics, Lower Silesian Center for Oncology, Pulmonology and Hematology, Poland