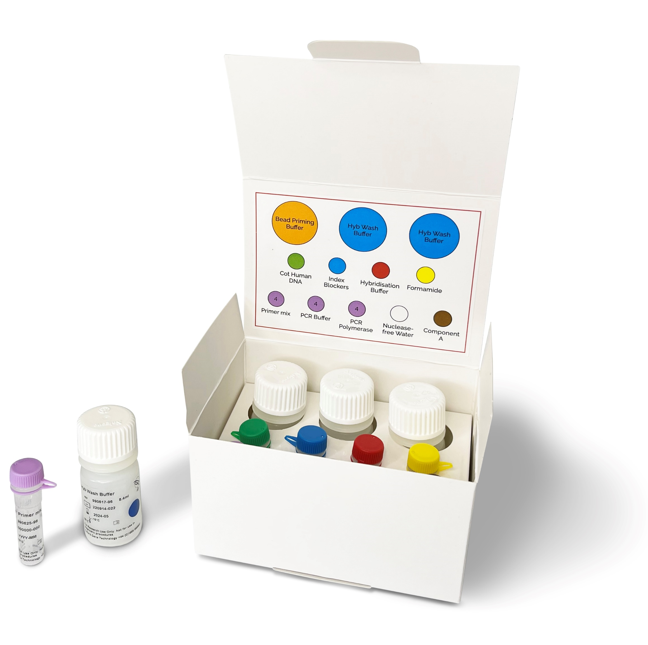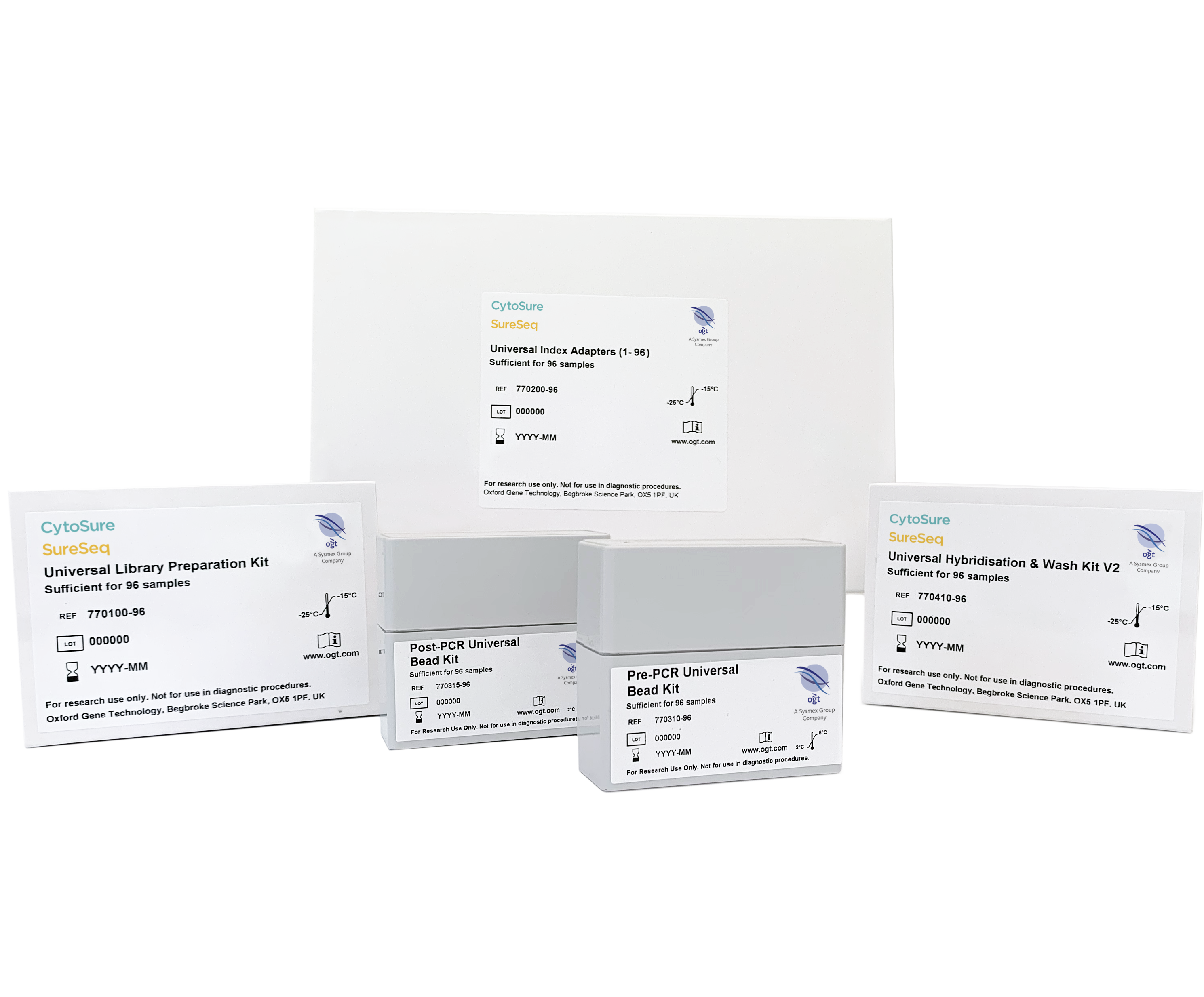
Library preparation protocols usually consist of lengthy multistep processes that require costly reagents and substantial hands-on time. We have listened to our customers’ feedback and developed the Universal NGS Workflow Solution V2, our latest and most advanced system for capture of targeted genomic regions and generation of NGS libraries.
OGT’s Universal NGS Workflow Solution V2, tested and optimised with both SureSeq™ and CytoSure® NGS panels, includes new combined multi-enzymatic fragmentation, end repair and A-tailing step. Together, with convenient bead concentration steps, the kit delivers increased convenience and flexibility for our highest quality library preparation.
The workflow has fewer clean up and QC steps, delivering scalability and reproducibility while minimising human errors. Additionally, the inclusion of Unique Dual Index (UDI)/Unique Molecular Index (UMI) adapters in the first phase of library preparation increases the kit multiplexing efficiency and confidence, enhancing capabilities to include sensitive applications. The included Universal Hybridisation & Wash Kit simplifies this key step while offering excellent coverage uniformity and reproducibility.

A simplified system including all necessary reagents, without the need for expensive supporting hardware; assuring a guaranteed output

New enzymatic fragmentation, end repair and A-tailing improves sample throughput

Inclusion of UDI and UMI adapters increases multiplexing efficiency and confidence, delivering robust and reliable results

Choice of pack size, with 24 or 96 unique dual indexes delivering multiplexing efficiency
Universal NGS Workflow Solution V2 is our latest and most advanced system for capture of targeted genomic regions and generation of NGS libraries, tested and optimised with both SureSeq™ and CytoSure® NGS panels.
OGT’s Universal NGS Workflow Solution V2 has a combined multi-enzymatic fragmentation, end repair and A-tailing step followed by convenient bead concentration steps, leading to the need of fewer clean-up and QC steps, to deliver scalability and reproducibility while minimising human errors.
Additionally, the inclusion of Unique Dual Index (UDI)/Unique Molecular Index (UMI) adapters in the first phase of library preparation increases the kit multiplexing efficiency and confidence, enhancing capabilities to include sensitive applications. OGT’s Universal Hybridisation & Wash Kit simplifies this key step while offering excellent coverage uniformity and reproducibility.
The Universal NGS Workflow Solution V2 uses a Unique Dual Indexing strategy to ensure accurate demultiplexing and avoid index hopping and Unique Molecular Identifier (UMI) for reliable identification of low-frequency variants.
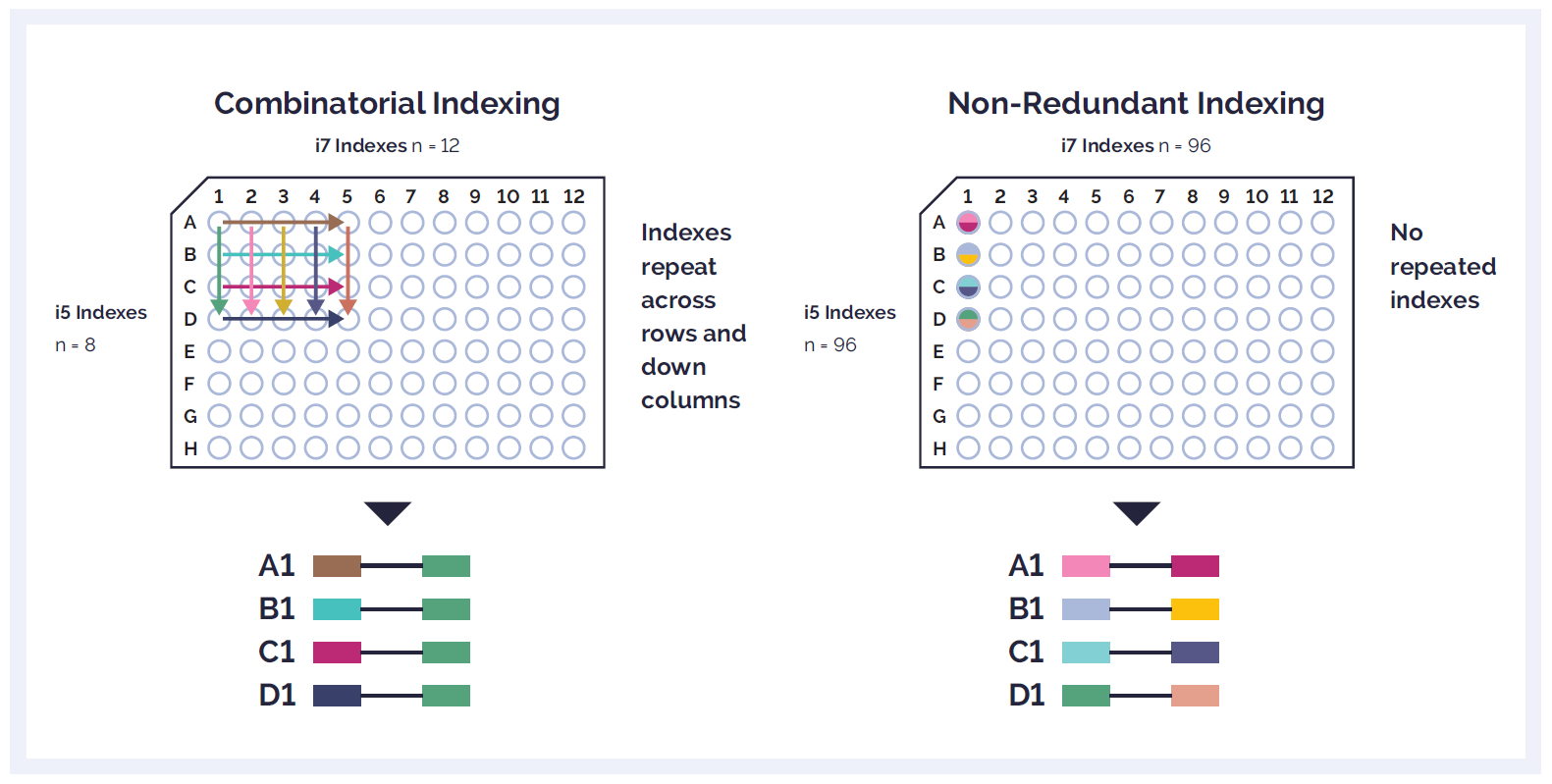
Figure 2: Combinatorial dual indexing barcoding (A) uses a plate matrix approach of i5 and i7 adapters that leads to unique combinations of non-unique indexes. In contrast, Unique Dual Indexing (B) uses completely unique indexes across the whole plate, avoiding any repetition in sequence (eg, 96 unique i5s and 96 unique i7s per 96-well plate).

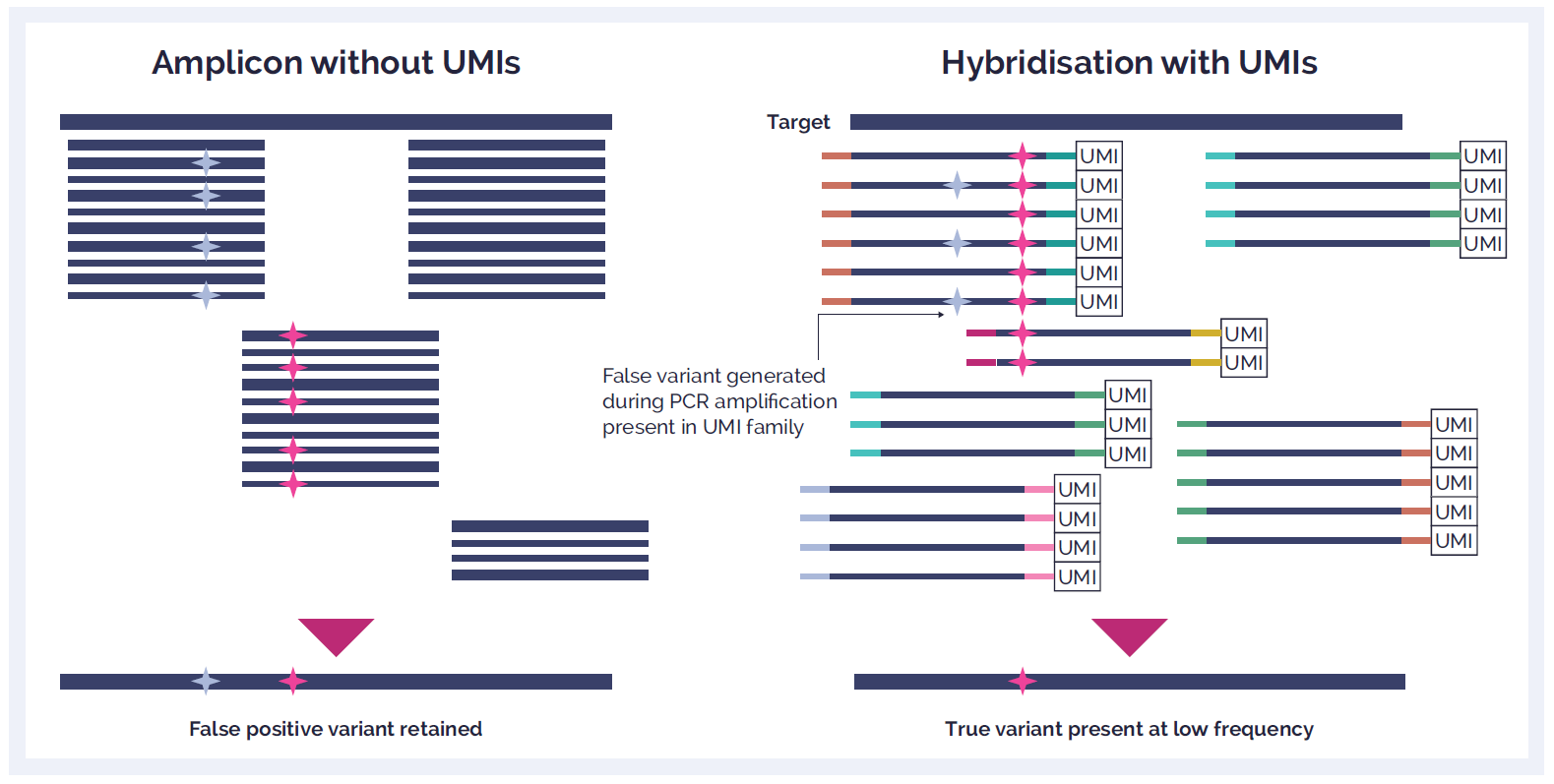
Figure 3: UMIs are short arbitrary oligonucleotides sequences that are attached to the library of DNA fragments by ligation prior to the amplification step. Reads that have the same UMI tag are from the same original DNA fragment and so the deriving PCR amplicons should be identical. Without the use of UMIs, low-frequency variants can be confused with DNA polymerase errors produced during the amplification step as well as sequencing errors produced during the sequencing step.

The Universal NGS Workflow Solution V2 works hand-in-hand with both CytoSure NGS Constitutional kit as well as with SureSeq targeted cancer enrichment panels, ensuring you get the most sensitive and reproducible variant detection and industry-leading coverage uniformity.
Interpret is OGT’s powerful, easy-to-use and customisable next-generation sequencing analysis solution delivers accurate calling of SNVs and indels, as well as structural aberrations, including ITDs, PTDs, CNVs, LOH and translocations.
Interpret is designed to work seamlessly with SureSeq and CytoSure NGS panels offering flexible accessibility for data analysis; whether through a stand-alone computer, laboratory server or another web-enabled device.
OGT partnership approach is key to providing the highest level of service, working closely with you to understand your unique challenges, customising our approach to meet your exact needs.
Discover our bespoke solutions with customised NGS Panel and software customisation options.