Cutaneous melanoma (CM) is the most dangerous form of skin tumour and causes 90% of skin cancer mortality1. With recurrent somatic mutations in BRAF, NRAS, KIT and NF1 among the most common genetic aberrations underlying pathogenesis of melanoma, next generation sequencing (NGS) has been an invaluable tool in helping to characterise the overall genomic landscape of melanomas.
Choose your ideal melanoma NGS panel from our range of fully tested and optimised NGS panel content. Simply mix and match the genes or individual exons you require and get the most out of your sequencing runs. Use in conjunction with the SureSeq™ FFPE DNA Repair Mix† for improved NGS library yields, %OTR and mean target coverage from challenging FFPE derived samples.

Detect low frequency melanoma variants consistently with confidence

Create your ideal panel and sequence only what’s relevant for your research

Get the most comprehensive insight into disease-driving mutations

Easy-to-use analysis solution for accurate detection of all variants
The most frequently activated pathway in melanoma is the mitogen-activated protein kinase (MAPK) pathway, often activated through mutations in the V600 codon of BRAF (in 35–50% of melanomas) and the Q61 codon of NRAS (10–25%)2, with mutations being mutually exclusive.
Mutations of KIT are found in particular subsets of melanoma, where the mutations activate signal-transduction pathways (MAPK and PI3K) that ultimately lead to cell proliferation. Approximately 70% of KIT mutations identified in melanoma are found in exon 11, most commonly L576P (Figure 1).
Neurofibromatosis type 1 (NF1) is a relatively common tumour predisposition syndrome related to germline aberrations of NF1, a tumour suppressor gene‡. Recent studies have additionally shown NF1 to play a critical role in somatic events in a wide range of tumours, including melanoma. The tumour suppressor function of neurofibromin is largely attributed to a small central region which comprises 360 amino acids encoded by exons 20-27a3. OGT’s expert bait design offers excellent uniformity for all of these key genes associated with melanoma (Figure 2).
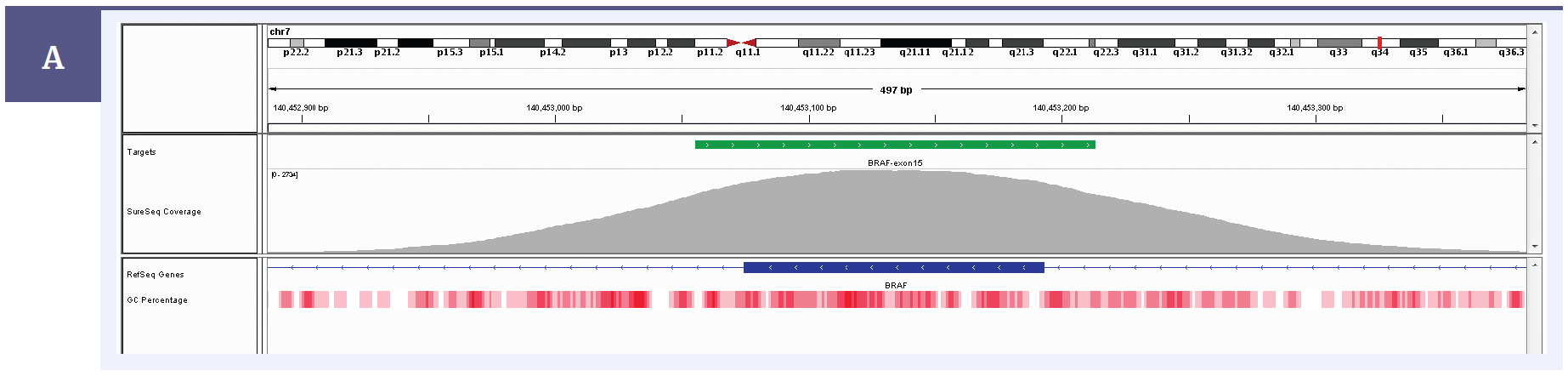
Figure 1a: Illustration of the exceptional uniformity of coverage of BRAF exon 15 with a SureSeq melanoma panel. Depth of coverage per base (grey). Targeted region (green). Gene coding region as defined by RefSeq (blue). GC percentage (red).

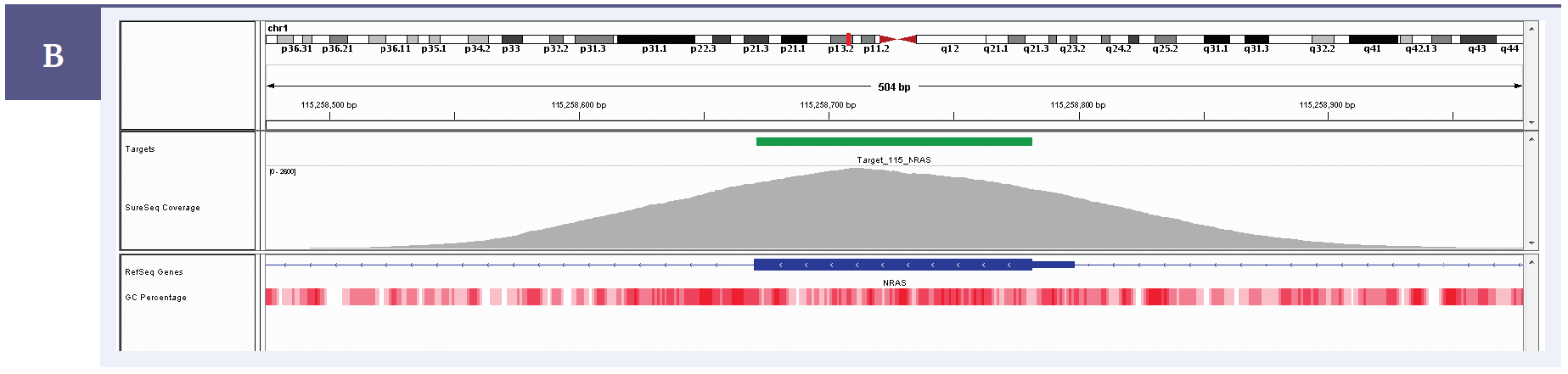
Figure 1b: Illustration of the exceptional uniformity of coverage of NRAS exon 2 with a SureSeq melanoma panel. Depth of coverage per base (grey). Targeted region (green). Gene coding region as defined by RefSeq (blue). GC percentage (red).

* Exon examples not yet available

*Panel size is an estimation based upon recommended custom panel content. †The SureSeq FFPE DNA Repair Mix can only be purchased in conjunction with SureSeq NGS panels, not as a standalone product. ‡Due to the presence of pseudogenes in NF1, it is recommended that an orthogonal technique is used to verify any mutations detected.