Lyudmila Georgieva, Stephanie Carpenter, Natalie Milner, Venu Pullabhatla, James Reid, Kay Wiebrands, Aysel Heckel, Graham Speight
Loss-of-function mutations in tumor suppressor genes BRCA1 and BRCA2 have been implicated in an increased risk for breast and ovarian cancer1,2. Screening for germline mutations in these genes allows research into familial risk of developing breast and ovarian cancer. In addition, assessment of somatic mutations in tumor samples can help research into tumor development, drug response and the development of new therapies.
A wide range of genetic variations are associated with breast and ovarian cancer, including single nucleotide variants (SNVs), small insertions/deletions (indels) and copy-number variations (CNVs). For more than a decade, the gold standard for mutational screening has been Sanger sequencing and multiplex ligation-dependent probe amplification (MLPA), imposing significant time and cost burden.
Advances in next-generation sequencing (NGS) now allows for the reliable detection of CNVs in addition to SNVs/indels in a single assay.
In this study, we tested the capability of the SureSeq™ Breast Cancer + CNV Panel to overcome the challenges currently experienced and provide a possible future single assay to be developed for breast and ovarian cancer.
The SureSeq hybridization-based approach was used to assess the detection of CNVs as well as SNVs and indels in a single assay; the workflow is outlined in Figure 1.
The SureSeq Breast Cancer + CNV Panel can be used for detection of SNV/indels and CNVs in 7 genes - ATM, BRCA1, BRCA2, TP53, CHEK2, PALB2, PTEN.
A number of different sample types were profiled:
The resulting libraries were sequenced using a 2x150 bp read length (v2) protocol on an Illumina MiSeq®.
CNV detection concordance was assessed by comparing NGS calls to events reported by an orthogonal technology, MLPA for set 1 samples and CytoSure® Cancer +SNP Arrays and CytoSure software (OGT) for set 2 samples.
Data sequencing analysis including CNV detection was performed using Interpret, OGT’s complimentary NGS analysis software.
Facilitated by OGT’s expert bait design, the hybridization-based SureSeq Breast Cancer + CNV Panel delivers excellent coverage uniformity, allowing consistent detection of SNVs and indels in germline samples, as well as somatic analysis on FPPE tissues down to 1% VAF (Figures 2 and 3).
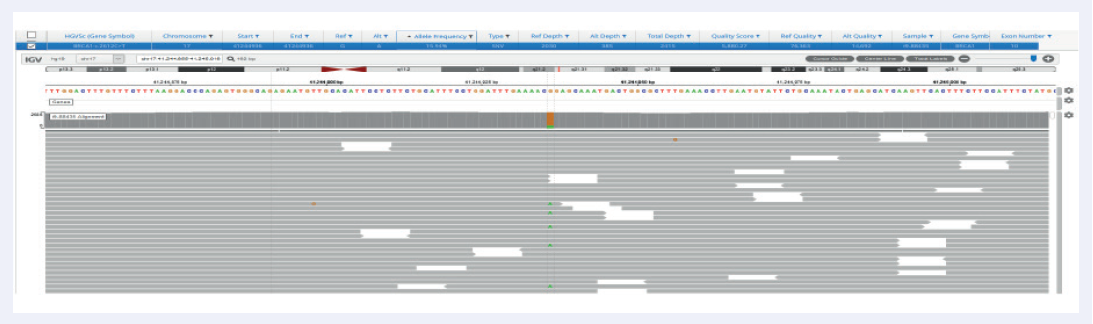
Figure 3: Excellent uniformity and high depth of coverage allowing confident detection of variants. Example of BRCA1 exon 10 missense variant Pro871Leu (rs799917) with frequency 15.9% identified in FFPE samples. Data generated with OGT SureSeq protocol averaging ~2000x deduplicated coverage. Depth of coverage per base (grey).

SureSeq and CytoSure: For research use only. Not for diagnostic procedures
Call +44 (0)1865 856800 Email contact@ogt.com
Send us a message and we will get back to you